Research
Genomics and extinction risk
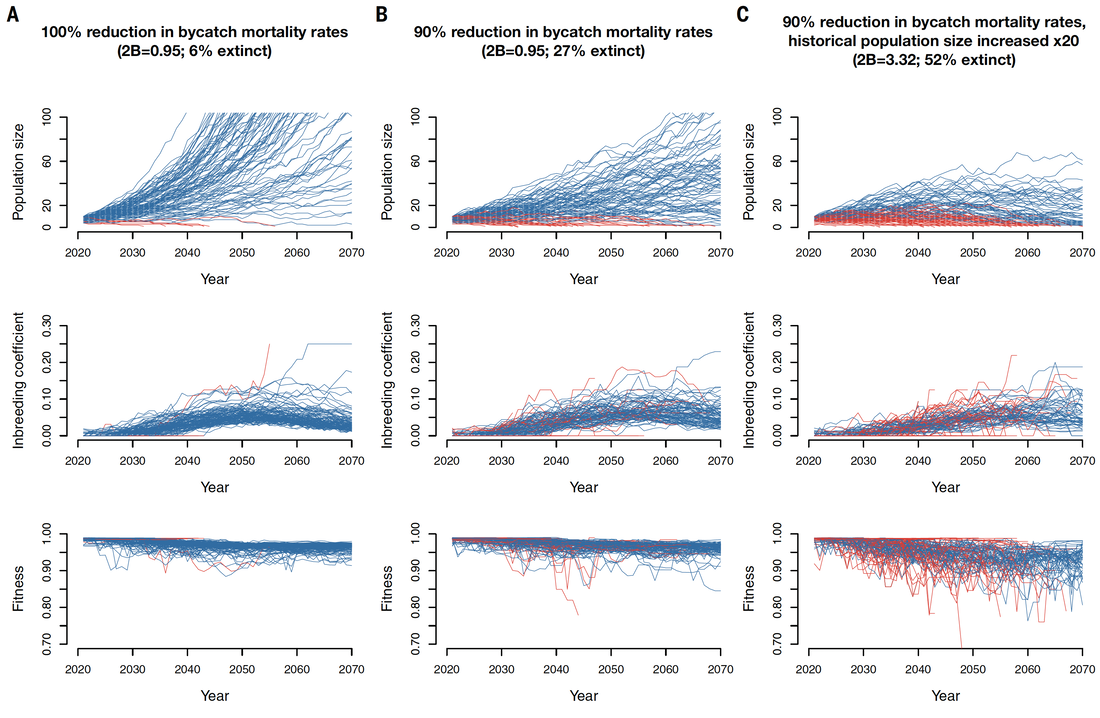
My primary research interest is understanding the ways in which genetic factors can contribute to extinction in small wildlife populations. Much of my work has focused on using eco-evolutionary models in SLiM to explore the role of demographic history in determining levels of recessive strongly deleterious variation in a population, and detailing how such mutations can greatly contribute to extinction risk when exposed by inbreeding . I have also employed such models in concert with genomic datasets to investigate the dynamics of deleterious variation and extinction risk in species including the vaquita porpoise, the Isle Royale moose population, and the California sea otter population. In addition, I have contributed to several recent review papers aiming to synthesize progress in this field and encourage broader adoption of computational simulation tools in conservation genomics.
Inferring selection and demography parameters from genomic datasets
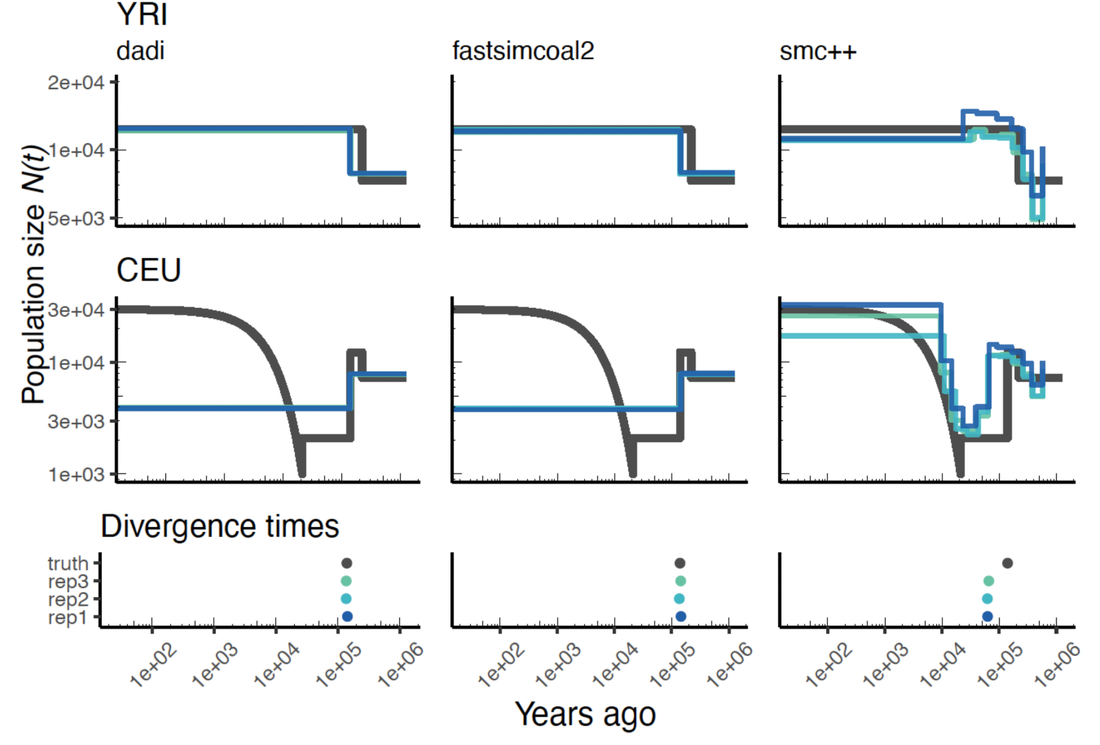
A central aim in population genetics is to infer demography and selection parameters. Quantifying these parameters provides fundamental insights into the interplay between selection and demography in natural populations, and are a key component of modelling population dynamics and extinction risk. Over the past several decades, much progress has been made in using genomic variation datasets to infer demography and selection parameters. However, many inference methods remain inadequately tested, and parameter estimates are rarely validated with orthogonal sources of information. My research seeks to better test existing inference methods and refine important parameter estimates. With the PopSim Consortium, I previously worked to test various popular methods for inferring demographic history from genomic data. More recently, my work has focused on refining estimates of selection and dominance parameters derived from the site frequency spectrum.